 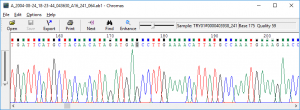
Since the app includes translation functions, you will be able to visualize the effect of a certain mutation with minimal effort. Uncomplicated DNA analysis tool with a user friendly design and amino-acid translation capabilitiesĤPeaks comes with a basic search tool which you can employ to quickly highlight a specific motif: you can use the predefined ones, or effortlessly make your own. At the same time, 4Peaks offers support for integrating user developed plug-ins that deal with specific tasks. The next step is to open the DNA sequence file you want to analyze: 4Peaks will generate a visual representation that can be easily edited by double clicking on the call you want to change.įor your convenience, 4Peaks comes with multiple translation dictionaries and BLAST algorithms: simply choose the ones you want to employ via the appropriate menus. Like with many other OS X software solutions, the 4Peaks installation process is reduced to dragging and dropping the app to your disk. Straightforward DNA sequence trace editor and viewer, with support for protein and DNA analysis plug-ins Moreover, although quite minimalist looking, the 4Peaks utility is able to handle most sequence trace file formats, such as SCF, ZTR, EXP, PLN, BIO, ALF, CTF, and a lot more. Samtools sort 8 -O bam -o sample1.tmp.4Peaks is a comprehensive Mac app designed to provide uncomplicated yet highly efficient tools for viewing and analyzing information extracted while sequencing DNA samples. 1ĪlignmentSieve -numberOfProcessors 8 -ATACshift -bam sample1.bam -o Or, we could do this using alignmentSieve. # use -X 2000 to allow larger fragment size (default is 500)īowtie2 -very-sensitive -X 2000 -x $Bowtie2Index -1 $ ' > fragments.bed # better alignment results are frequently achieved with -very-sensitive I will use Bowtie2 (since it was used in many tutorals and papers). Two popular aligners are BWA and Bowtie2. The next step is to align reads to a reference genome. Just like analyzing other NGS data, quality control is needed for raw FASTQ, and there are many programs avaliable, such as Trimmomatic and Trim Galore. Some steps are optional, like merging BAMs. The following pipeline includes several common analysis in ATAC-seq setting, from data trimming to peak calling. Ferhat Ay Lab - ATAC-seq processing pipeline.Parker Lab - ATAC-seq lab for BIOINF525.Tobias Rausch - ATAC-seq analysis pipeline.ENCODE - ATAC-seq Data Standards and Prototype Processing Pipeline. 
Harvard FAS Informatics - ATAC-seq Guidelines – clear and up-to-date.use as few PCR cycles as possible when constructing the library.each replicate has 25 million non-duplicate, non-mitochondrial aligned reads for single-end sequencing and 50 million for paired-ended sequencing.See Buenrostro et al., 2015, ENCODE - ATAC-seq Data Standards and Prototype Processing Pipeline, and Harvard FAS Informatics - ATAC-seq Guidelines for details. generate occupancy profiles of TFs (footprinting).identify important transcription factors.map accessible chromatin across tissues or conditions.It utilizes a hyperactive Tn5 transposase to insert sequencing adapters into open chromatin regions.ĪTAC-seq overview ( Buenrostro et al., 2015).Īnd the peaks look like the following figure. ATAC-seq overviewĪTAC-seq (Assay for Transposase-Accessible Chromatin with high-throughput sequencing) is a method for determining chromatin accessibility across the genome. I could not indicate all the sources, but I want to thank them for sharing experiences/knowledge/code. The content were compiled from multiple resources on the internet (forum, papers, workshop etc.). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
